Overcome Cell Viability Measurement Challenges
Flow cytometry and manual hemocytometry have long been the go-to methods for measuring cell viability and apoptosis. But their drawbacks—including clogging issues, sample wastage, labor intensity and slow speeds—can impede progress in critical research or production processes.
Distinguishing between cellular material from non-cellular particles is also difficult. Even with fluidics-based techniques, there is little contrast to help identify subvisible particles, impacting data accuracy along the way.
Because of this, it has been hard to find a fast, reliable, and high-throughput approach that can accurately measure cell viability and purity—but Aura CL is here to change that.
Determine the Success of Your Cell Therapy with Aura CL
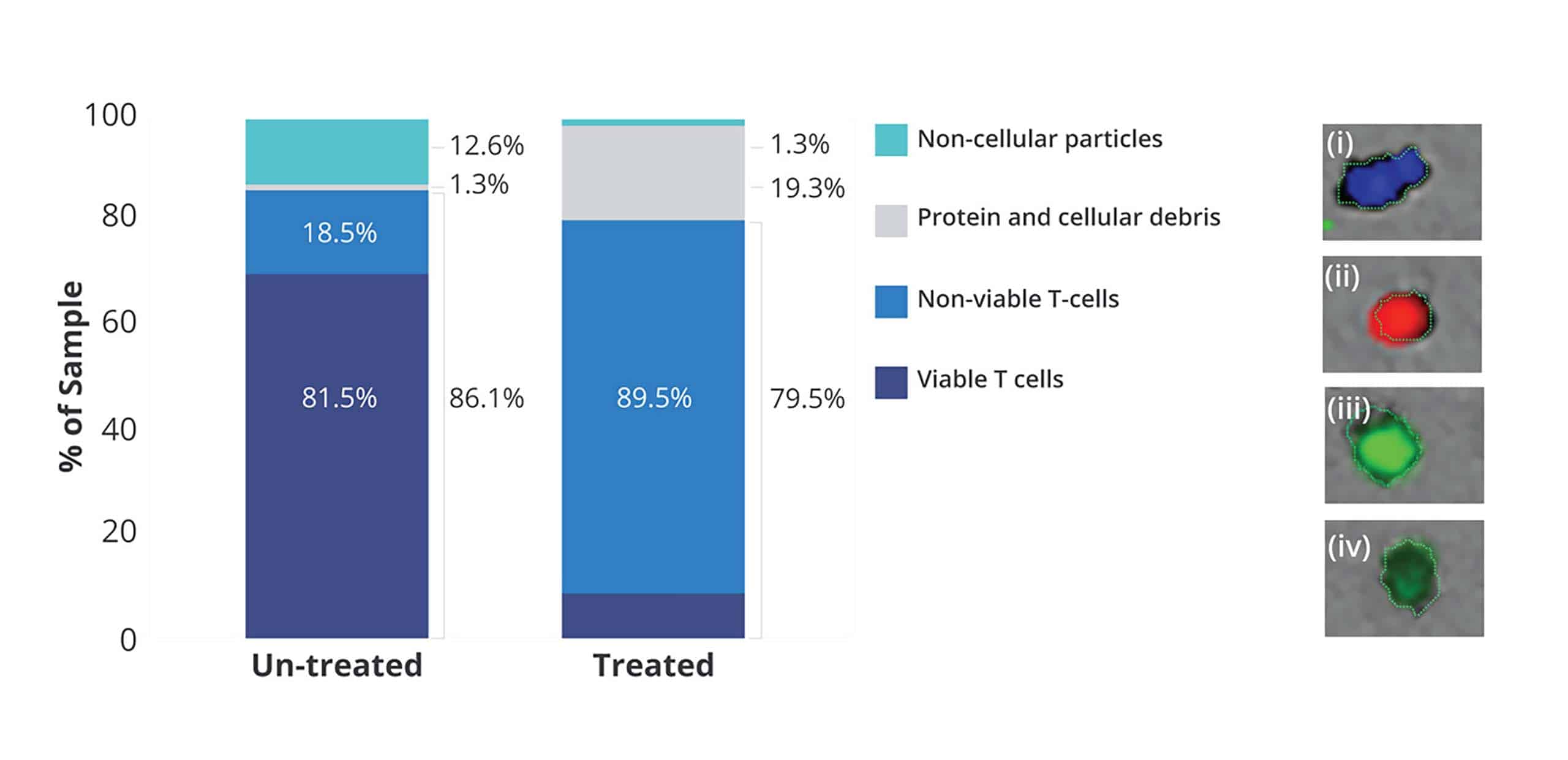
With Aura CL, cell therapies are rigorously examined for quality and safety, ensuring they meet the specifications outlined by USP 1046.
Powered by Backgrounded Membrane Imaging (BMI) and Fluorescent Membrane Microscopy (FMM), this game-changing technology provides reliable indications of viable cells in your cell therapy product.
Add in Particle Vue software, and you get a speedy analysis on 96-well formats - allowing precise measurement of not only viability but also purity. Making sure that whether it's iPSCs or T-cells, you have all the information necessary with just one assay.
Identify Non-Viable and Viable Cells in Your Therapeutic
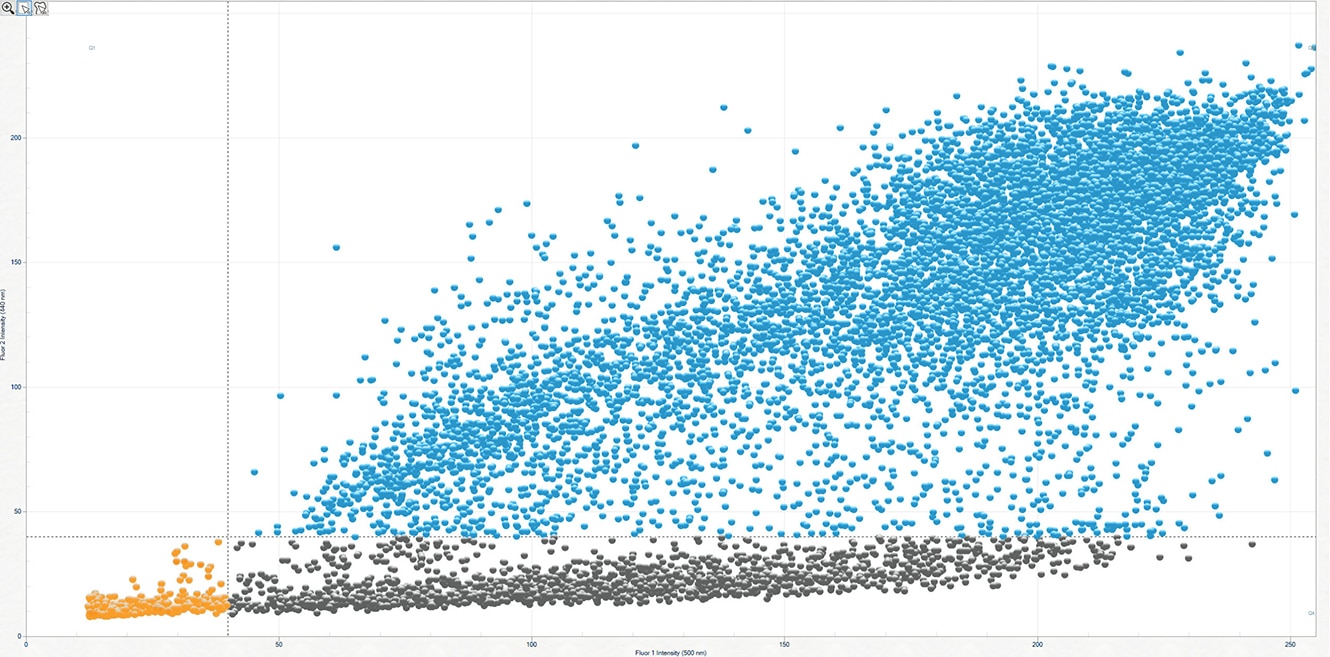
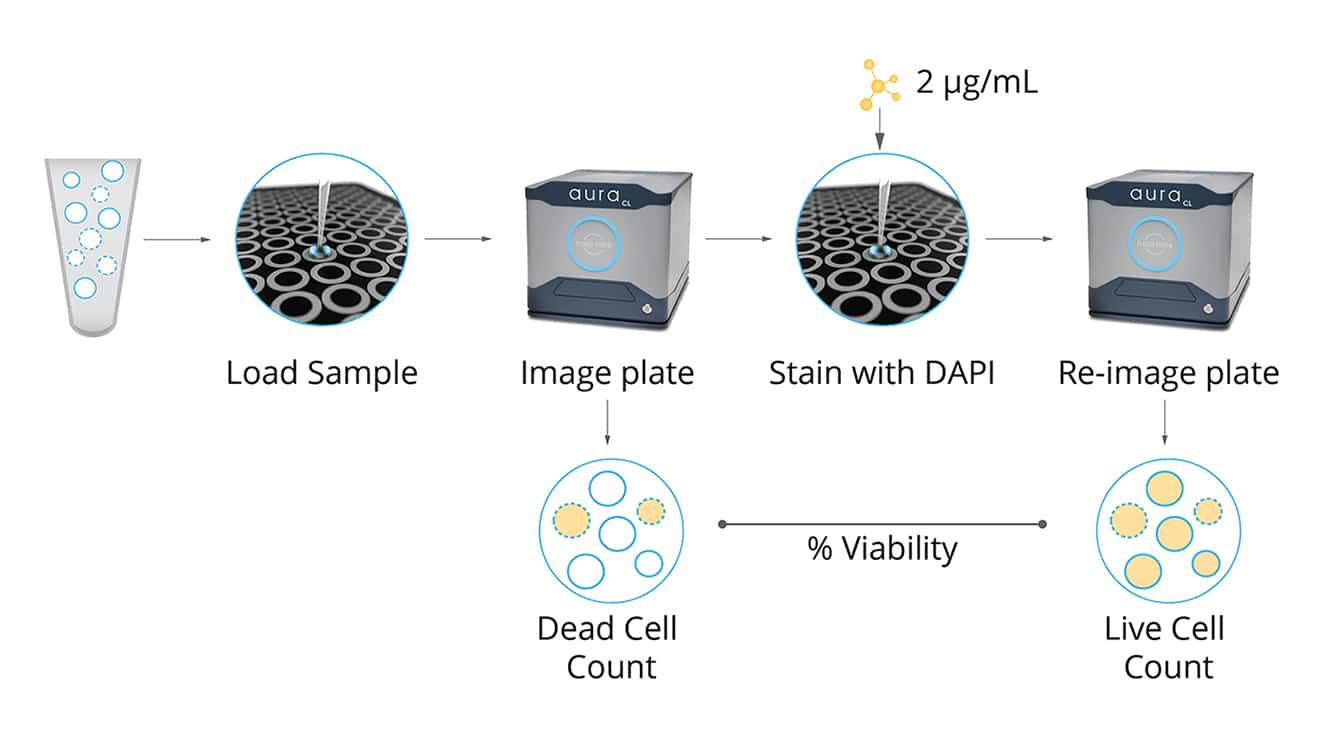
Aura CL's cell viability assay takes advantage of the increases in plasma membrane permeability observed during apoptosis to accurately identify and measure viable and non-viable cells.
After samples are incubated with a low concentration of a cell viability stain such as DAPI, only non-viable cells can be visualized due to the dye's impermeability. Utilizing Aura CL imaging technology, these labeled non-viable cells can be precisely quantified before applying an increased dose of the same staining agent in order to estimate and determine the amount of viable cell concentrations within each sample—producing results that rival traditional methods such as hemocytometry or flow cytometry.
Simplify Particle ID with FMM
Ensuring product quality and safety is paramount in the manufacturing process. However, using morphology alone to effectively identify particles with different root causes can be unreliable.
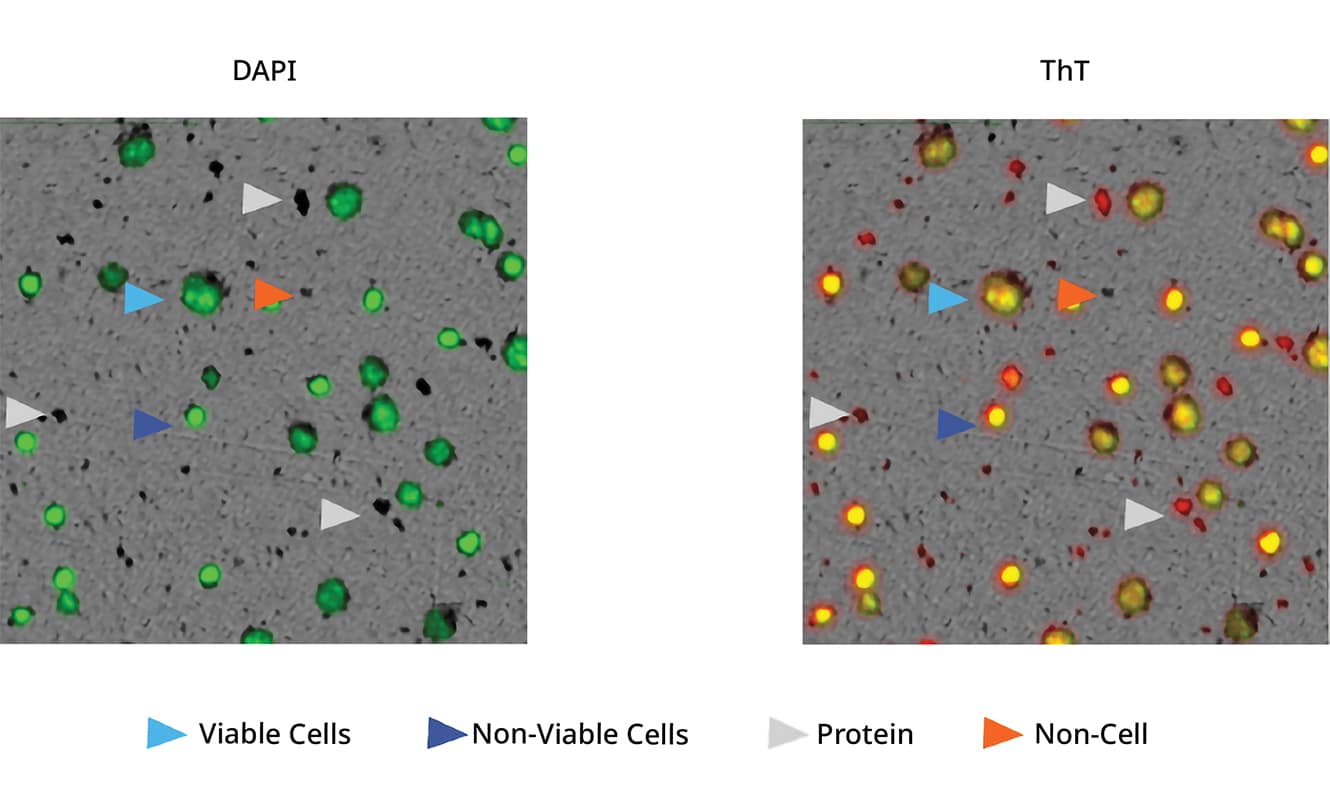
This is where FMM comes in. FMM offers an accurate solution by utilizing particle-specific dyes, like Thioflavin T (ThT), that bind to protein aggregates and simplify ID.
With FMM, you are able to easily distinguish between viable cells, non-viable cells, protein aggregates, and more – allowing for a reliable and efficient manufacturing process.
Featured products
Frequently Asked Questions
Cell viability testing is essential for evaluating the health and functionality of cells under specific conditions. It serves as a quality control measure, ensuring reliable and reproducible results by identifying viable cells for experiments. In drug discovery, it helps assess the efficacy and safety of potential drug candidates, while also optimizing experimental conditions for cell growth and function. Moreover, cell viability assays enable predictive modeling of cellular responses, guiding future experiments and understanding disease progression. Overall, cell viability testing is fundamental for advancing research across various fields by providing insights into cellular behavior and facilitating the development of new therapies.
Fluorescence-based cell viability assays present distinct advantages for researchers. Their high sensitivity and specificity allows for the detection of subtle changes in cell viability, ensuring precise assessment of treatment effects. These assays offer quantitative analysis, facilitating accurate measurement and comparison of experimental conditions or drug responses. Their multiplexing capability enables simultaneous measurement of multiple parameters, providing comprehensive insights into cellular behavior. Fluorescence assays are adaptable to various cell types and experimental setups, making them versatile tools across different research fields. Additionally, their compatibility with automation streamlines workflows, enhancing efficiency and reproducibility.
Determining the "best" cell viability assay depends on various factors, such as your specific research objectives, the characteristics of the cells being studied, and available resources. However, among the widely used assays, fluorescence-based methods are often favored for their high sensitivity, quantitative analysis, and versatility. Popular fluorescence-based assays include the MTT assay, resazurin assay, and the CellTiter-Glo assay. Each of these assays has its strengths and limitations, and the choice depends on factors such as sensitivity requirements, throughput, and compatibility with experimental conditions.
The cell viability parameter is the measure of the percentage of live and functional cells within a population. It serves as an indicator of the overall health and viability of the cell population under specific experimental conditions. Cell viability can be assessed using various assays that measure aspects of cellular health, such as metabolic activity, membrane integrity, or cellular ATP levels. Evaluating cell viability is crucial in research, particularly in drug discovery, toxicology, and basic cell biology, as it provides insights into the effects of treatments, environmental factors, or genetic manipulations on cellular physiology and function.